A Complete Guide to Single-Cell VDJ Sequencing Data Analysis
📋 Overview • 📁 File Preparation • 🚀 Main Pipeline • 📊 Results Interpretation
This document provides a detailed guide for analyzing single-cell VDJ sequencing data using dnbc4tools.
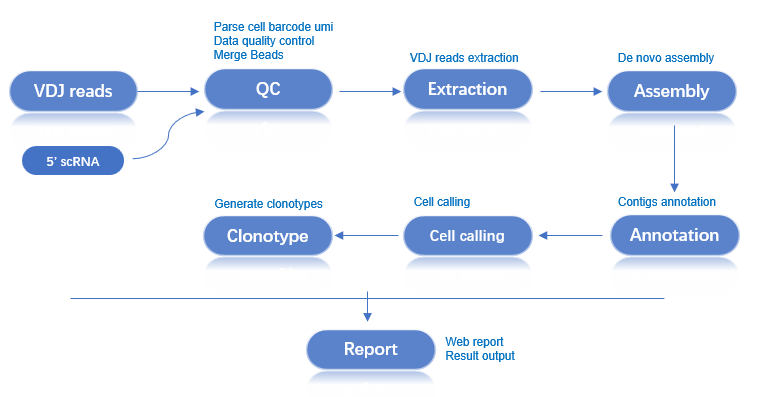
Workflow: 5' Transcriptome Analysis → VDJ Library Processing → Sequence Assembly & Annotation → Cell Filtering → Clonotype Analysis → Analysis Report

For transcriptome analysis, please refer to dnbc4tools rna run. The 5' transcriptome analysis pipeline requires adding the --end5 parameter to the standard single-cell RNA analysis workflow.
Here is an example script to generate an expression matrix for a single sample:
$dnbc4tools rna run \
--name sample_5rna \
--cDNAfastq1 /data/sample_cDNA_R1.fastq.gz \
--cDNAfastq2 /data/sample_cDNA_R2.fastq.gz \
--oligofastq1 /data/sample_oligo1_1.fq.gz,/data/sample_oligo2_1.fq.gz \
--oligofastq2 /data/sample_oligo1_2.fq.gz,/data/sample_oligo2_2.fq.gz \
--genomeDir /opt/database/Homo_sapiens \
--threads 30 \
--end5
The analysis requires the following files:
| File Type | Description |
|---|---|
| FASTQ Files | VDJ library sequencing data containing TCR or BCR sequence information. |
| singlecell.csv File | The cell information file from the 5' transcriptome analysis results. |
The analysis requires the singlecell.csv file from the 5' transcriptome analysis output directory. This file contains merged information from the cell and barcode columns, as well as an is_cell_barcode column (1 for a cell, 0 for a non-cell), which is used to identify valid cells.
Example file content:
cell,reads,gene,umi,is_cell_barcode,barcode
CELL1118_N3,1485813,5693,57580,1,AGATCGCCTACGATCACGAT;GGTGGAAGGTGAGAGAAGCG;GTAGTTCTAGGCTAAGTACT
CELL1651_N3,805447,4881,32131,1,ATCTCAAGCCCACCGTGTGT;CATCAATTAAGTGATCGCAT;CCTAACTGAGGAACGCTTAG
CELL4_N3,820269,5649,30326,1,AACACCTGATCGTTCAATAA;AATTCGAAGGTAGTCGGAAT;CACATGTTACATGTTCTATA
CELL906_N3,656624,4853,28142,1,ACGTCCGCGTGACCATGTGC;AGGAGCTCCATTGATCTTAA;CAATCCGGAGAACGTATCTG
CELL1577_N2,672637,4610,24822,1,ATATTCTCACGTAACGGATG;TAGGAACTCGGCTTAGATCT
CELL2064_N2,542608,2355,23324,1,CATAAGCACTCACCGCTAGT;GTCTCACAGTTGTTCACTAG
CELL1332_N6,580950,4702,22953,1,AGGTGTAAGCCTACCGGACC;CGCACTCACCTAACATTGTG;CTTGCCGCGCTATCAATGCA;GCCACTAGTCGACGCGGTTG;GTCAGCATGCTCTTCCACAG;TCGATATCCTCACTCTTAAC
CELL726_N4,585934,4617,22660,1,ACCTACGGCGTTACTATGTG;CGACGCTCTCGACAGTTAGG;CGGCAGAGTCTTGGCGCTTA;TCCGACCGTATCTTCATCTC
CELL4010_N1,555308,4268,22554,1,AGAGAGTCGCAGCAAGCGAC
The main VDJ pipeline combines single-cell VDJ library data with the 5' transcriptome results from the same sample. It includes these key steps:
To run TCR analysis for a single sample, use the following example script:
$dnbc4tools vdj run \
--name sample_tcr \
--fastq1 /data/sample_tcr_R1.fastq.gz \
--fastq2 /data/sample_tcr_R2.fastq.gz \
--ref human \
--chain TR \
--beadstrans /sample_5rna/outs/singlecell.csv \
--threads 10
To run BCR analysis for a single sample, use the following example script:
$dnbc4tools vdj run \
--name sample_bcr \
--fastq1 /data/sample_bcr_R1.fastq.gz \
--fastq2 /data/sample_bcr_R2.fastq.gz \
--ref human \
--chain IG \
--beadstrans /sample_5rna/outs/singlecell.csv \
--threads 10
After auto-detecting the dark reaction, the software begins the analysis. Here is an example:
2025-04-23 23:01:01 Performing VDJ data processing
Chemistry(darkreaction) determined in fastqR1: darkreaction
2025-04-23 23:01:02 Processing VDJ library filtering...
...done
2025-04-23 23:31:13 Preparing data for VDJ assembly...
...done
2025-04-24 00:21:14 Sequence Assembly and annotation of VDJ Gene Segments
...done
2025-04-24 02:49:16 Cell calling for VDJ analysis...
...done
2025-04-24 02:51:52 Generating VDJ clonotype analysis...
...done
2025-04-24 02:53:49 Converting VDJ results from cellbarcode to cellid...
...done
2025-04-24 02:53:58 Statistical analysis and report generation for results.
...done
Analysis Finished Elapsed Time: 3:53:07
A successful run will end with Analysis Finished.
Upon completion, outs (outputs) and logs directories will be generated. The outs directory includes:
.
├── airr_annotations.tsv
├── all_contig_annotations.csv
├── all_contig.fasta
├── all_contig.fasta.fai
├── *_scVDJ_IG_report.html
├── clonotypes.csv
├── consensus_annotations.csv
├── consensus.fasta
├── consensus.fasta.fai
├── filtered_contig_annotations.csv
├── filtered_contig.fasta
├── filtered_contig.fasta.fai
└── metrics_summary.xls
Related Documentation:
Content to be added